Installation
You can install the package from CRAN:
install.packages("edibble")You can install the development version with:
# install.packages("remotes")
remotes::install_github("emitanaka/edibble")Overview
The goal of edibble R-package is to aid in the plan, design and simulation of experiments by mapping fundamental components of experiments to an object oriented system. The edibble system is built on the principle that the system must make it easy to recover experimental context by encouraging the user to be explicit about experimental details in fundamental terms.
Examples
Consider an experiment where you want to know what is an effective way of teaching (flipped or traditional style) for teaching a particular subject and how different forms of exams (take-home, open-book or closed-book) affect student’s marks.
There are four classes for this subject with each class holding 30 students. The teaching style can only be applied to the whole class but exam can be different for individual students.
library(edibble)
set.seed(2020)
des <- design(name = "Effective teaching") %>%
set_units(class = 4,
student = nested_in(class, 30)) %>%
set_trts(style = c("flipped", "traditional"),
exam = c("take-home", "open-book", "closed-book")) %>%
allot_trts(style ~ class,
exam ~ student) %>%
assign_trts("random")
serve_table(des)
#> # An edibble: 120 x 4
#> class student style exam
#> <U(4)> <U(120)> <T(2)> <T(3)>
#> <chr> <chr> <chr> <chr>
#> 1 class1 student001 traditional take-home
#> 2 class1 student002 traditional closed-book
#> 3 class1 student003 traditional open-book
#> 4 class1 student004 traditional take-home
#> 5 class1 student005 traditional closed-book
#> 6 class1 student006 traditional take-home
#> 7 class1 student007 traditional take-home
#> 8 class1 student008 traditional open-book
#> 9 class1 student009 traditional take-home
#> 10 class1 student010 traditional closed-book
#> # ℹ 110 more rowsBefore constructing the experiment, you might want to think about what you are recording for which level of unit and what values these variables can be recorded as.
out <- des %>%
set_rcrds_of(student = c("exam_mark",
"quiz1_mark",
"quiz2_mark",
"gender"),
class = c("room",
"teacher")) %>%
expect_rcrds(exam_mark <= 100,
exam_mark >= 0,
quiz1_mark <= 15L,
quiz1_mark >= 0L,
quiz2_mark <= 30L,
quiz2_mark >= 0L,
factor(gender, levels = c("female", "male", "non-binary", "unknown"))) %>%
serve_table()
out
#> # An edibble: 120 x 10
#> class student style exam exam_mark quiz1_mark quiz2_mark gender
#> <U(4)> <U(120)> <T(2)> <T(3)> <R(120)> <R(120)> <R(120)> <R(120)>
#> <chr> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 class1 student001 traditio… take-ho… o o o o
#> 2 class1 student002 traditio… closed-… o o o o
#> 3 class1 student003 traditio… open-bo… o o o o
#> 4 class1 student004 traditio… take-ho… o o o o
#> 5 class1 student005 traditio… closed-… o o o o
#> 6 class1 student006 traditio… take-ho… o o o o
#> 7 class1 student007 traditio… take-ho… o o o o
#> 8 class1 student008 traditio… open-bo… o o o o
#> 9 class1 student009 traditio… take-ho… o o o o
#> 10 class1 student010 traditio… closed-… o o o o
#> # ℹ 110 more rows
#> # ℹ 2 more variables: room <R(4)>, teacher <R(4)>When you export the above edibble design using the export_design function, the variables you are recording are constraint to the values you expect, e.g. for factors, the cells have a drop-down menu to select from possible values.
export_design(out, file = "/PATH/TO/FILE.xlsx")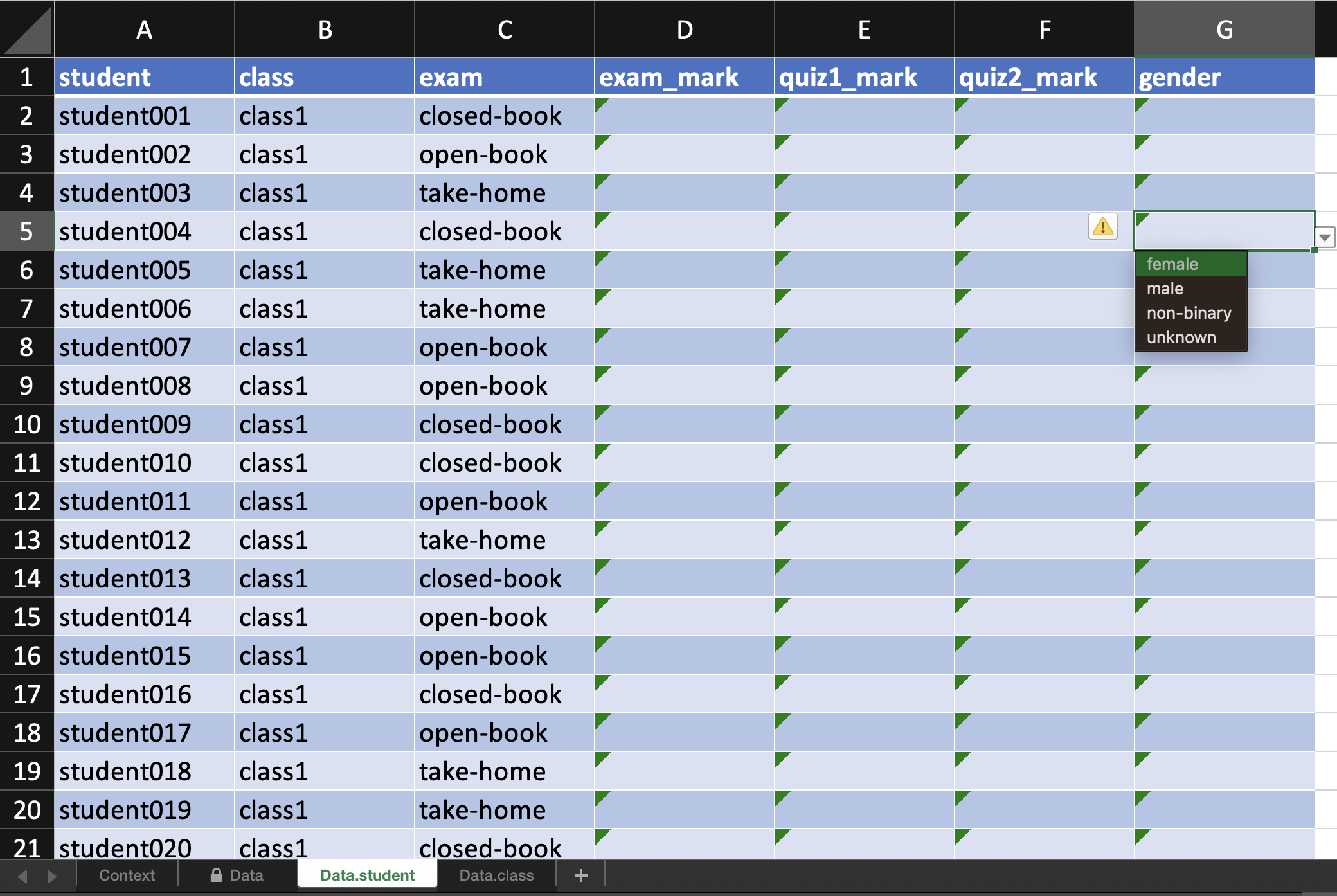
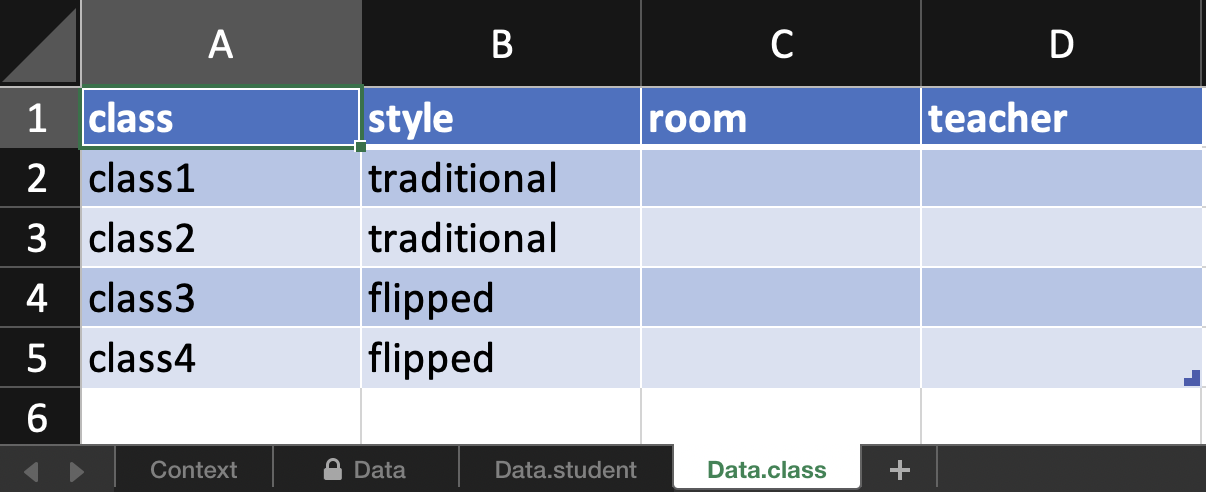
In addition, there is a spreadsheet for every observational level. E.g. here room and teacher is the same for all students in one class so rather than entering duplicate information, these are exported to another sheet for data entry.
There is also support for more complex nesting structures. You can always make the structure using edibble and take the resulting data frame to use in other experimental design software. It’s also possible to bring existing data frame into edibble if you want to take advantage of the exporting feature in edibble.
design("nesting structure") %>%
# there are 3 sites labelled A, B, C
set_units(site = c("A", "B", "C"),
# each site has 2 blocks except B with 3 sites
block = nested_in(site,
"B" ~ 3,
. ~ 2),
# levels can be specified by their number instead
# so for below "block1" has 30 plots,
# "block2" and "block3" has 40 plots,
# the rest of blocks have 20 plots.
plot = nested_in(block,
1 ~ 30,
c(2, 3) ~ 40,
. ~ 20)) %>%
serve_table()
#> # nesting structure
#> # An edibble: 190 x 3
#> site block plot
#> <U(3)> <U(7)> <U(190)>
#> <chr> <chr> <chr>
#> 1 A block1 plot001
#> 2 A block1 plot002
#> 3 A block1 plot003
#> 4 A block1 plot004
#> 5 A block1 plot005
#> 6 A block1 plot006
#> 7 A block1 plot007
#> 8 A block1 plot008
#> 9 A block1 plot009
#> 10 A block1 plot010
#> # ℹ 180 more rowsExperimental data
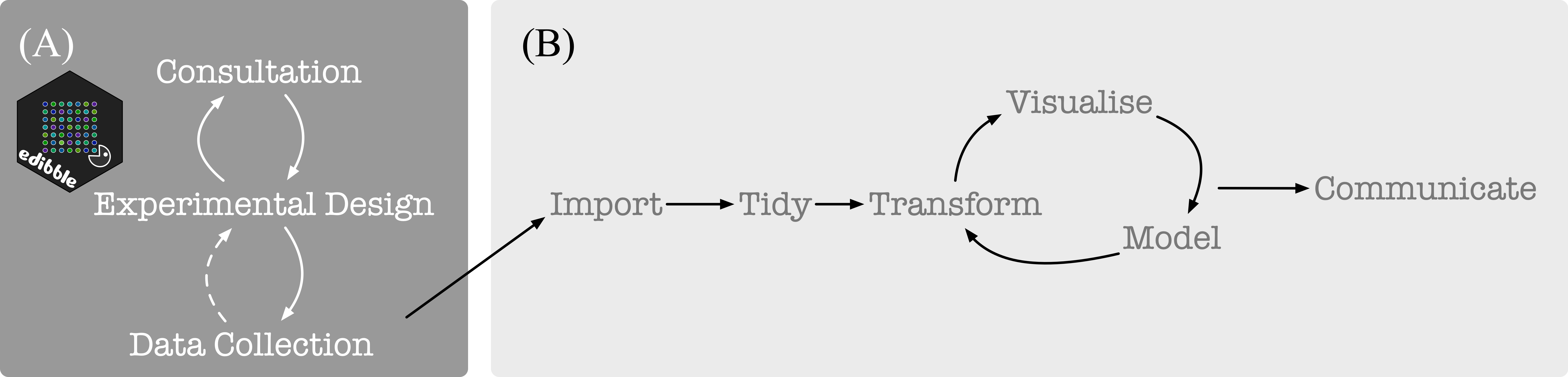
tidyverse is well suited for the data science project workflow as illustrated below in (B) (from Grolemund and Wickham 2017). For experimental data, the statistical aspect begins before obtaining data as depicted below in (A). The focus of edibble is to facilitate work in (A).
The edibble R-package differ considerably to other packages for constructing experimental design with a focus on the whole process and less on the randomisation process (which the other software generally focus and do well on). Some features include:
- declaratively create experimental designs based on experimental components (e.g. units and treatments),
- explicitly specify variables that are to be recorded (e.g. response), and
- set expected values for variables to be recorded which restrict the data entry when the design is exported as an xlsx file,
- simulate values for record variables,
- make classical named designs see Cookbook chapter.
Limitations
Currently, edibble:
- expects you to know the number of units available from the start. Unknown numbers will be supported in future versions.
- edibble supports experiments that are not comparative experiments.
- does not do enough testing so design should be diagnosed after construction (which should be done regardless of how much testing edibble implements).
Related Work
The way that edibble specifies experimental design is largely novel (if I say so myself) and there are no work that resembles it. I’m concurrently working on two extension packages:
-
deggust- to visualise the designs constructed from edibble as ggplot2 objects (WIP). -
sizzled- for experiments that require sample size calculation (WIP).
Below are some other related work. You can also have a look at the CRAN Task View for Design of Experiment and Analysis of Experimental Data for a whole collection.
-
DeclareDesignfor survey or sampling designs -
designrfor balanced factorial designs with crossed and nested random and fixed effect to data frame -
daefor functions useful in the design and ANOVA of experiments (this is in fact powering the randomisation in edibble) -
plotdesignrfor designing agronomic field experiments
Acknowledgement
edibble is hugely inspired by the work of Tidyverse Team. I’m grateful for the dedication and work by the Tidyverse Team, as well as R Development Core Team that supports the core R ecosystem, that made developing this package possible.
Tidyverse familiarity
The implementation in edibble adopt a similar nomenclature and design philosophy as tidyverse (and where it does not, it’s likely my shortcoming) so that tidyverse users can leverage their familiarity of the tidyverse language when using edibble. Specifically, edibble follows the philosophy:
- main functions do one thing and have a consistent form of
<verb>_<noun>(e.g.set_unitsandset_rcrds) where the nouns are generally plural. Exceptions are when the subject matter is clearly singular (e.g.designandset_context); - pipable functions;
- all dots arguments are dynamic dots;
- duplicate names repaired with same option as
tibblefor additions to edibble graph; - ability for developers to extend certain components.
- the specification of complex nested structure drawing similarity to
dplyr::case_when(LHS is character or integer for edibble however).
Code of Conduct
Please note that the edibble project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.
